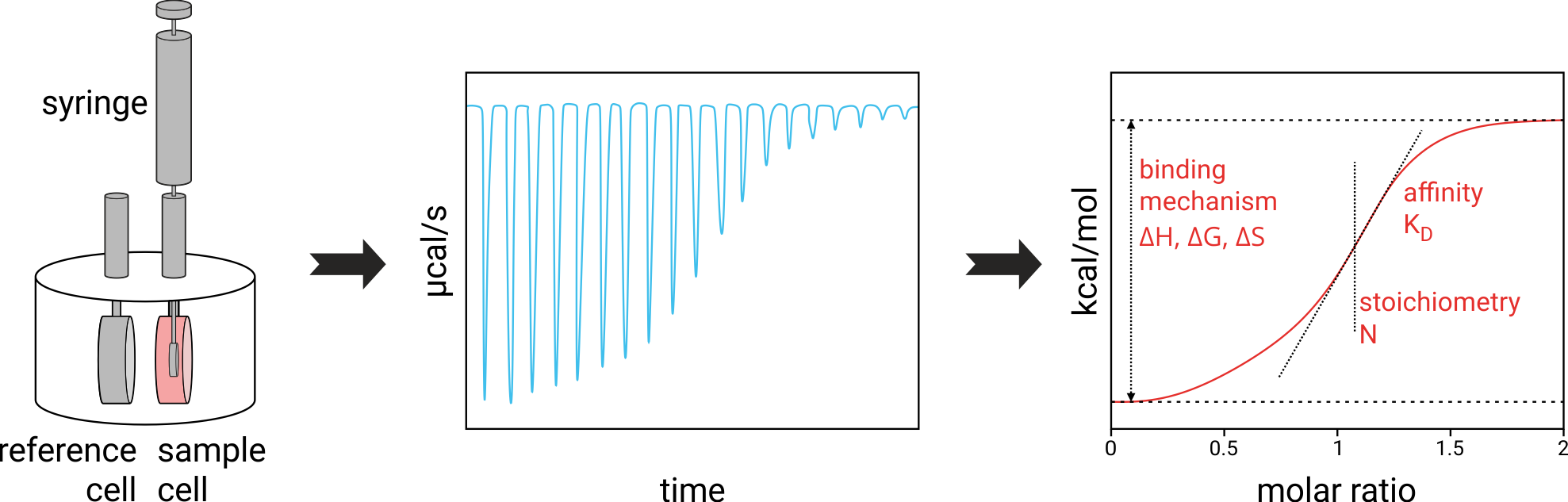
Molecular interactions that can be measured with ITC
ITC works with almost all classes of target and ligand molecules, as long as they are available in the required volumens (typically 60-300 µL) and concentrations (typically 1-50 µM for proteins). In ITC, the target molecule is loaded into a 300 µL volume cell and the ligand molecule is titrated via an automatic syringe.
If you are uncertain whether your type of interaction can be measured with ITC or if you want to use the 2bind ITC service just leave us a message:
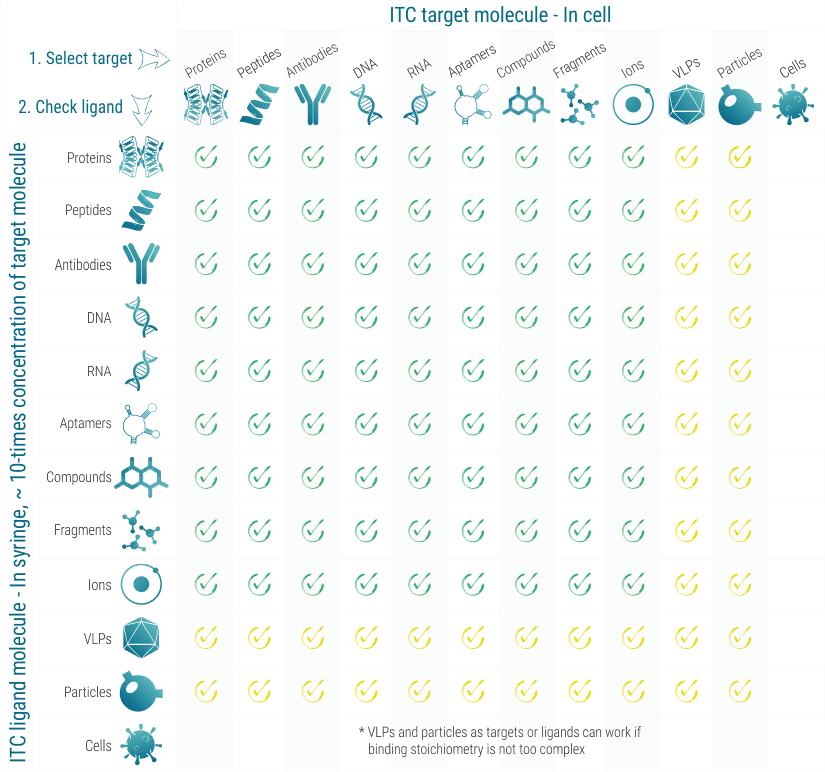
What ITC can tell
Binding thermodynamics
ITC excels in revealing the enthalpy or binding heat (ΔH) and entropy (ΔS) of molecular interactions as well as giving the free enthalpy of binding (ΔG).
Steady-State Affinity
ITC can tell the affinity of molecular interactions (dissociation constant KD) derived from the thermodynamic measurement.
Interaction stoichiometry
ITC will directly tell you the precise stoichiometry (i.e. 1:1 or 2:1 binding) of your molecular interaction.
- In-depth characterization of molecular interactions
- Inhibitor or lead-candidate ranking in drug discovery
- Protein-protein interaction analysis
- Drug discovery (compound-based and fragment-based)
- Protein characterization
- Additive screening
- Enzyme-inhibitor screening
- Complex binding analysis
- Mode-of-action analysis
- Protein complex characterization
- Analysis of binding mechanisms
2bind ITC Services
2bind offers a variety of services using the Isothermal Titration Calorimetry technology. Benefit from comprehensive and precise analysis of molecular interactions. Enjoy thermodynamic profiling or your interaction of interest.
The 2bind ITC services can be combined as you choose for all questions in drug discovery, antibody development, protein biophysics and analysis, as well as aptamer characterization.
ITC – Technology and FAQs
Overview
ITC is widely used for quantifying binding affinity, for drug development (candidate optimization and validation), for measuring thermodynamics of molecular interactions, for confirmation of binding targets in small-molecule drug-discovery, for determination of binding specificity, or for validation of IC50 and EC50 values.
Technology
ITC directly measures the heat released or absorbed during molecular binding events and the concomitant formation of molecular complexes. The label-free, in-solution characteristic of ITC experiments allows for the direct and native-like determination of all important parameters that characterize the thermodynamics of a molecular interaction: the binding constant (KD), the reaction stoichiometry (n), the observed binding enthalpy (ΔHobs), the observed binding entropy (ΔSobs), the observed heat capacity of binding (∆Cobs) and finally the change in free enthalpy of binding (ΔG). Thus, ITC generates a complete thermodynamic profile of a molecular interaction and, for example, can help to differentiate between binding reactions that are driven by enthalpy (due to the formation of non-covalent interactions across the binding-interface) or by entropy (due to the release of water molecules from a binding pocket).
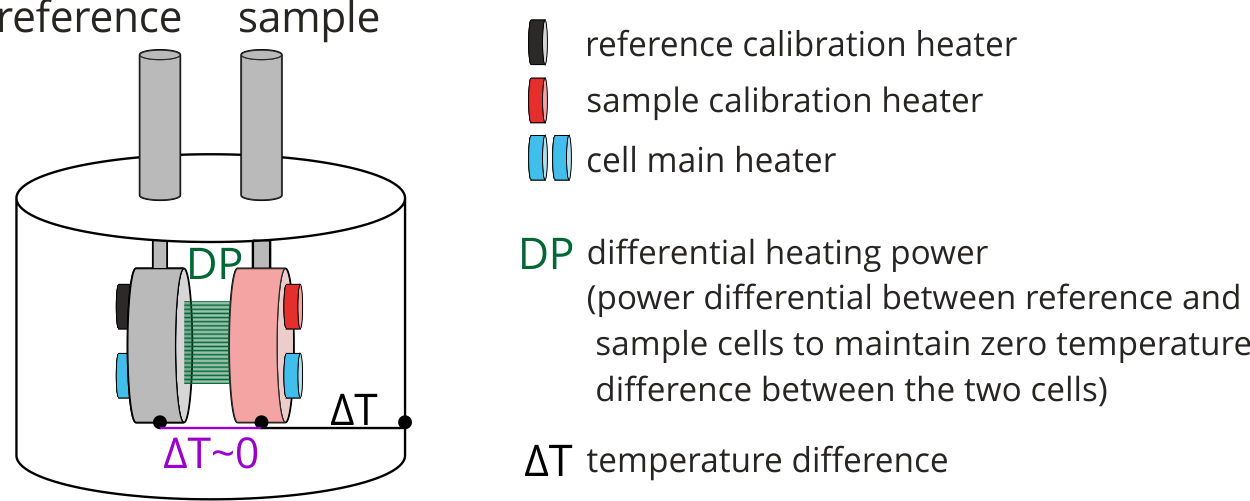
In order to determine the heat released or absorbed during molecular binding events an ITC apparatus comprises two coin-shaped cells: A reference cell, which is usually filled with water and a sample cell, which contains one of the two interaction partners. The two cells are kept at the same constant temperature so that the temperature difference (ΔT) is zero. Heat changes due to binding events in the sample cell are registered by the device by measuring the differential power (DP) that is required to maintain a temperature difference of zero between the sample and the reference cell. For example, in the case of an exothermic binding event, heat is released inside the sample cell and the heating power required to keep the sample cell at the predefined fixed temperature is lower than that for the reference cell. The other interaction partner (often called the ligand) is titrated into the sample cell via a rotating syringe, typically in aliquots of 0.5 to 2 µL. Each injection leads to a heat change in the sample cell, which is recorded as a heat pulse (see Figure 2). The heat pulses are integrated over time to generate a titration curve that related the heat (kcal/mol of injected ligand) to the molar ratio (amount of titrated interaction partner relative to amount of interaction partner in the sample cell). Fitting the resulting isotherm to different binding models (e.g. 1:1 binding) yields the thermodynamic parameters described above.
Here at 2bind, the state-of-the-art Malvern PEAQ ITC platform is used. This system features exceptional sensitivity and excellent reproducibility. The low sample consumption enables the analysis of interactions in cases where only small amounts of proteins or other macromolecules are available. Typically, the sample cell is filled with 200 µL of one interaction partner at a concentration of 10 to 30 µM and the syringe is filled with 40 µL of the other interaction partner at a concentration of 100 to 300 µM. The PEAQ-ITC systems allows a throughput of about 6 to 10 measurements per working day. Interactions can be quantified in the temperature range from 25 to 80 °C.
Typical applications
- Determining enthalpic and entropic contribution to molecular interactions (and derived the steady-state affinity)
- Determining changes in free energy upon molecular binding
- Determining stoichiometry of molecular interactions
- in various contexts:
protein-protein interactions
protein-DNA interactions
protein-RNA interactions
protein-small molecule interactions
DNA-small molecule interactions
RNA-small molecule interactions
For more information on potential applications take a look at our Application Database as well as the potential applications of ITC in Drug Discovery.
Advantages
- No labeling required → close-to-native conditions
- No immobilization required → measurements in solution
- Low sample consumption for ITC standards → only 300 µl of a 20 µM protein solution required
- Complete thermodynamic profiling → ΔH, ΔG, ΔS
- Wide temperature range → from 10°C to 80°C
- Tells binding characteristics → affinity and stoichiometry
- Wide range of binding affinities → range from nM to mM affinities measurable
- Wide molecule size range → small molecules to MDa
A comparison of ITC with other biophysical interaction analysis methods is available in our Technology Comparison Guide.
FAQ – General
What kind of molecular interaction can be measured?
Can I measure without labeling molecules?
Can I measure without immobilizing molecules?
What information do I get from an ITC experiment? Are thermodynamics possible?
Which range of affinities can be quantified?
Can I measure binding kinetics with ITC?
What is necessary for a successful determination of the thermodynamic parameters of an interaction?
FAQ – Samples
What are the required sample concentrations?
f these requirements are a concern, it is best to consider alternative methods like MicroScale Thermophoresis or nanoDSF that both require much lower sample concentrations (nM to µM range).
How much sample do I need?
Can I measure the binding of small molecules?
Do I need to prepare my samples in a specific way?
f these requirements are a concern, it is best to consider alternative methods like MicroScale Thermophoresis, nanoDSF, or Biolayer Interferometry that allow measurements in almost any kind of buffer and biological liquid.
FAQ – Assay Conditions
How high is the throughput?
For high-throughput analysis of molecular interactions it is best to consider MicroScale Thermophoresis or nanoDSF, as both give a much higher throughout compared to ITC. For more information on high-throughput interaction analysis take a look at our Drug Discovery Services.